Klebsiella pneumoniae in Germany: An overview on spatiotemporal distribution and resistance development in humans
Authors: Gamal Wareth, Lisa D. Sprague, Heinrich Neubauer and Mathias W. Pletz
Ger. J. Microbiol.
2021.
vol. 1, Iss. 1
pp:16-25
Doi: https://doi.org/10.51585/gjm.2021.0004
Abstract:
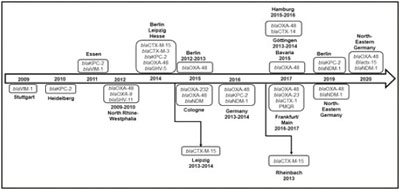
The emerging of multidrug-resistant Klebsiella pneumoniae (K. pneumoniae) is increasing worldwide. Rapid dissemination and increase of its incidence in Germany are observed and becoming a significant challenge for clinical laboratories and physicians. The current review highlights its chronological sequence of appearance and resistance development in humans in the past two decades in Germany. Emerging resistance problems of K. pneumoniae to the vast majority of available antimicrobial agents, including carbapenems and those of the ß-lactam group, were observed since the end of the last century and strains carrying diverse resistance patterns have emerged in most federal states of Germany. Still, several aspects of resistance development and pathogenesis are not fully understood. To date, hypervirulent K. pneumoniae (hvKp) isolates have been rarely isolated from German patients. The most frequent resistance genes identified are blaOXA-48, blaCTX-M-15, blaKPC-2, blaOXA-9, blaSHV-11, blaSHV-5 blaCTX-M-3, blaCTX-M-14, blaVIM-1 and the plasmid-encoded quinolone resistance (PMQR) gene. One Health genomic surveillance of K. pneumoniae strains from different reservoirs is required. This would help to understand in great detail the mechanisms leading to resistance development, spread and transmission, and developing alternative treatment regimens.
Keywords:
Klebsiella pneumoniae, Emerging, MDR, Review, Germany
Statistics:
Article Views: 3150
PDF Download: 80